Multi-omics for healthcare materials

The group Multi-omics for healthcare materials focuses on the mapping of disease-associated cellular signaling networks in order to inform precise imunoengineering solutions. Activities of the group are centered on generating and analyzing biomedical multi-omics datasets that can assist development of new materials for personalized healthcare applications.
Our research work aims to enable rational design of new materials by capitalizing on large-scale biomedical datasets. This involves development of new software tools, employment of advanced data analytics and modeling of material properties. Through collaborations with material scientists, we evaluate the efficiency of the in silico designed functionalized nanoparticles and other modified materials. Together with clinical and industrial partners, we seek to implement these new concepts into next generation healthcare solutions.
Group's background

Signaling in tumor microenvironment
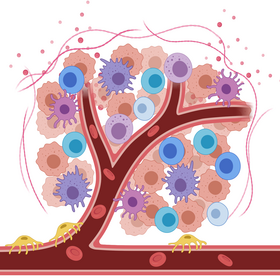
Signaling pathways in disease cells
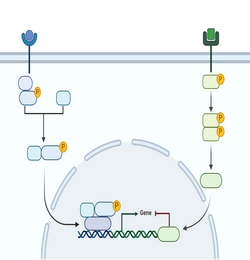
Nanoparticles as drug delivery systems
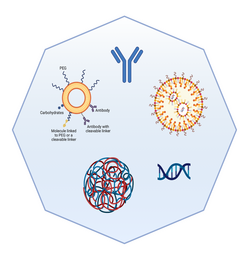
Lukas Häuser, PhD

Lukas.Haeuser@empa.ch
Alma Mater: Heidelberg University
Katharina Hast, PhD Student
Katharina.Hast@empa.ch
Alma Mater: Friedrich-Alexander-Universität Nürnberg-Erlangen
Jonas Bossart, PhD Student
Jonas.Bossart@empa.ch
Alma Mater: ETH
Tiberiu Totu, PhD Student
Tiberiu.Totu@empa.ch
Alma Mater: EPFL
Alumni
Benoit Hohl, MSc Student
Alma Mater: EPFL
Emile Bourban, MSc Student
Alma Mater: EPFL
Feifei Xia, MSc Student
feifei.xia@empa.ch
Alma Mater: UZH
Delia Cella, MSc Student
delia.cella@empa.ch
Alma Mater: ETH
T Totu#, J Bossart#, K Hast, C Li, M Rottmar, B Sobottka, G Yu, V Ayala-Nunez, M Buljan*: Delineation of signaling routes that underlie differences in macrophage phenotypic states. bioRxiv (2024)
M Buljan*, A Banaei-Esfahani, P Blattmann, F Meier-Abt, W Shao, O Vitek, H Tang, R Aebersold*: A computational framework for the inference of protein complex remodeling from whole-proteome measurements. Nature Methods 20(3):1523-1529. (2023)
M Buljan*, P Wick: Tailoring design of nanomaterials and systems to individualize patient treatments. Chimia 76(3):236-241. (2022)
M Buljan, R Ciuffa, A van Drogen, A Vichalkovski, M Mehnert, G Rosenberger, S Lee, M Varjosalo, L Espona Pernas, V Spegg, B Snijder, R Aebersold*, M Gstaiger*: Kinase Interaction Network Expands Functional and Disease Roles of Human Kinases. Molecular cell 79(3):504-520. (2020)
M Buljan, P Blattmann, R Aebersold*, M Boutros*: Systematic characterization of pan-cancer mutation clusters. Molecular systems biology 14(3): e7974. (2018)
JK White, AK Gerdin, NA Karp, E Ryder, M Buljan, JN Bussell, J Salisbury, S Clare, NJ Ingham, C Podrini, R Houghton, J Estabel, JR Bottomley, DG Melvin, D Sunter, NC Adams, L Baker, C Barnes, R Beveridge, E Cambridge, D Carragher, P Chana, K Clarke, Y Hooks, N Igosheva, C Ismail, H Jackson, L Kane, R Lacey, DT Lafont, M Lucas, S Maguire, K McGill, RE McIntyre, S Messager, L Mottram, L Mulderrig, S Pearson, HJ Protheroe, LA Roberson, G Salsbury, M Sanderson, D Sanger, C Shannon, PC Thompson, E Tuck, VE Vancollie, L Brackenbury, W Bushell, R Cook, P Dalvi, D Gleeson, B Habib, M Hardy, K Liakath-Ali, E Miklejewska, S Price, D Sethi, E Trenchard, D von Schiller, S Vyas, AP West, J Woodward, E Wynn, A Evans, D Gannon, M Griffiths, S Holroyd, V Iyer, C Kipp, M Lewis, W Li, D Oakley, D Richardson, D Smedley, C Agu, J Bryant, L Delaney, NI Gueorguieva, H Tharagonnet, AJ Townsend, D Biggs, E Brown, A Collinson, CE Dumeau, E Grau, S Harrison, James Harrison, Catherine E Ingle, Helen Kundi, Alla Madich, Danielle Mayhew, T Metcalf, S Newman, J Pass, L Pearson, H Reynolds, C Sinclair, H Wardle-Jones, M Woods, L Alexander, T Brown, F Flack, C Frost, N Griggs, S Hrnciarova, A Kirton, J McDermott, C Rogerson, G White, P Zielezinski, T DiTommaso, A Edwards, E Heath, MA Mahajan, B Yalcin, D Tannahill, DW Logan, DG MacArthur, J Flint, VB Mahajan, SH Tsang, I Smyth, FM Watt, WC Skarnes, G Dougan, DJ Adams, R Ramirez-Solis, A Bradley, KP Steel: Genome-wide generation and systematic phenotyping of knockout mice reveals new roles for many genes. Cell 154(2):452-464. (2013)
M Buljan*, G Chalancon, S Eustermann, GP Wagner, M Fuxreiter, A Bateman, MM Babu*: Tissue-specific splicing of disordered segments that embed binding motifs rewires protein interaction networks. Molecular cell 46(6): 871-883. (2012)
Regardless of open calls, if you are interested to join the lab as a Master student, a PhD student or a Postdoctoral fellow and are eligible for an independent fellowship, please get in touch!
Nov 2023
We are very happy that Lukas Häuser is starting his PhD in the group.
Sep 2023
We are chairing a session on the Dynamics of immune processes at the Basel Computational Biology conference.
Feb 2023
A very warm welcome to Katharina Hast who is starting her PhD at Empa!
Jun 2022
We are very grateful to Uniscientia foundation for supporting our research.
Oct 2021
Marija has joined the National Advisory Board of the Swiss Personalized Health Network.
Oct 2021
A very warm welcome to Jonas Bossart and Tiberiu Totu who are starting their PhD in the group!
Jan 2021
The group has now joined the Swiss Institute of Bioinformatics!
-
Share
